|
11/25/2023 0 Comments Rmarkdown shiny 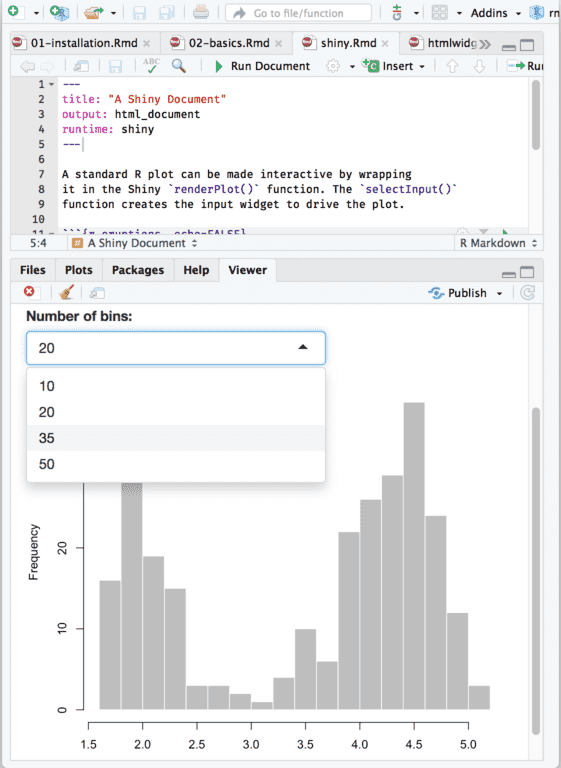
Mapping = aes(x = df_aggr_Result$hour, y = df_aggr_Result$`amount of samples`)) + Theme(plot.title = element_text(hjust = 0.5)) + Labs(title="amount of first-scan times over the day",x="hours", y = "amount of samples") + Geom_bar(stat="identity", boundary = 0, color="blue", fill="lightblue") + Mapping = aes(x = df_aggr_First_scan$hour, y = df_aggr_First_scan$`amount of samples`)) + I used following code to make a barplot of the amount of samples each hour of all the samples in r markdown: Another example is show the histogram, representing the amount of sample each hour of $test IGF. eg: show the histogram, representing the amount of sample each hour for $Source(location) A. Now the data is kind of prepared I would like to make a histogram/barplot to give the amount of samples over time (1 hour interval) and to use a dropdown menu to select an extra filter. I hope this give the same result on the Shiny app. #The difference between the ResultTime and the FirstScanTime is the turn around time:ĭf1$TS_start <- paste(df1$FirstScanDate, df1$FirstScanTime)ĭf1$TS_end <- paste(df1$ResultDate, df1$ResultTime)ĭf1$TAT <- difftime(df1$TS_end,df1$TS_start,units = "mins") #Create a subset of the data to remove/exclude the unnecessary columns: Server_data <- lim(file="PostCheckAnalysisRoche.txt", header = TRUE, na.strings=c(""," ","NA")) Setwd("G:/My Drive/Traineeship Advanced Bachelor of Bioinformatics 2022/Internship 2022-2023/internship documents/") #' #' The \code tags $ div ( tags $ head ( tags $ script ( src = "rmd_resources/rmd_loader.js" ), tags $ link ( href = "rmd_resources/rmd_loader.I use the code: #First the exported file from the Infinity server needs to be in the correct folder and read in: 
#' Run a Shiny document #' #' Start a Shiny server for the given document, and render it for display. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
